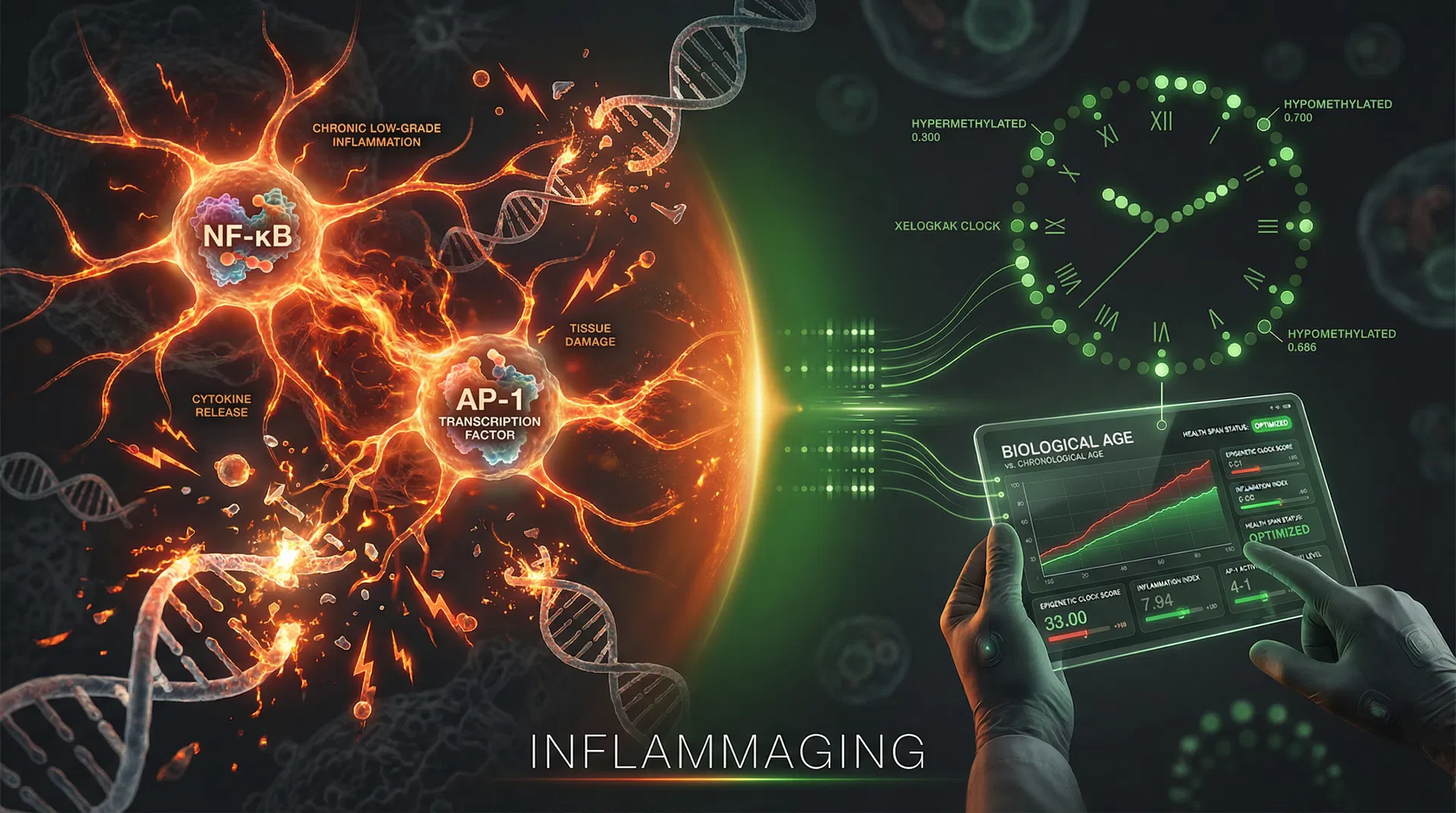
Inflammaging — the chronic, low-grade, sterile inflammation that accumulates with biological age — is not a side effect of ageing. It is an epigenetically encoded programme, driven by AP-1 transcription factor hijacking of the chromatin landscape, and it is the primary molecular mechanism through which epigenetic dysregulation converts into clinical disease. XELGEN's 850,000 CpG methylation platform is designed to read this programme at clinical resolution — detecting it years before symptoms appear, quantifying it with precision, and tracking its reversal under intervention.
Introduction: The Downstream Output of All Four Pillars
In Part I of this series (When the Epigenome Loses Its Memory), we described how ageing is driven by systems-level epigenetic fidelity failure across four interacting pillars: nuclear architecture deterioration, epigenetic memory collapse, nucleosome remodelling drift, and AP-1 transcriptional hijacking. In Part II (Why Single Biomarkers Fail), we demonstrated that no single biomarker can capture this complexity — that multi-factor epigenomic interaction explains up to 15× more variance in biological age than any individual exposure alone.
Part III addresses what happens when all four pillars fail simultaneously: the cell enters a state of chronic, low-grade, sterile inflammation that does not resolve. This state — inflammaging — is now understood to be the primary molecular mechanism through which epigenetic dysregulation converts into clinical disease, accelerated biological ageing, and organ system deterioration.
The critical insight from recent science is that inflammaging is not merely a consequence of ageing. It is an epigenetically encoded programme — one that can be measured, tracked, and, with the right intervention, reversed. XELGEN's 850,000 CpG methylation platform is specifically designed to read this programme at clinical resolution.
What Is Inflammaging?
The term was coined by immunologist Claudio Franceschi in 2000 to describe a paradox: the immune system of older individuals is simultaneously overactivated (producing chronic pro-inflammatory cytokines) and underperforming (failing to mount effective responses to pathogens and tumour cells). This dual dysfunction — chronic background fire with impaired targeted response — is now recognised as a unifying mechanism across virtually every age-related disease.
"Inflammaging is a highly significant risk factor for both morbidity and mortality in the elderly people, as most if not all age-related diseases share an inflammatory pathogenesis." — Franceschi & Campisi, Journal of Gerontology, 2014
The key inflammatory mediators in inflammaging include:
| Mediator | Source | Primary Effect |
|---|---|---|
| IL-6 | Senescent cells, adipose tissue | Systemic inflammation, muscle wasting |
| TNF-α | Macrophages, senescent fibroblasts | NF-κB activation, insulin resistance |
| IL-1β | NLRP3 inflammasome | Vascular inflammation, neuroinflammation |
| CRP | Liver (IL-6 driven) | Cardiovascular risk marker |
| CXCL10 | Interferon-stimulated cells | T-cell recruitment, autoimmune activation |
| MCP-1/CCL2 | Endothelium, macrophages | Monocyte recruitment, atherosclerosis |
What distinguishes inflammaging from acute inflammation is its chronicity and sterility — it is not triggered by infection or injury, and it does not resolve. The source is internal: senescent cells secreting the senescence-associated secretory phenotype (SASP), mitochondrial DNA leaking into the cytoplasm and activating cGAS-STING, and — critically — AP-1 transcription factor hijacking of the epigenome.
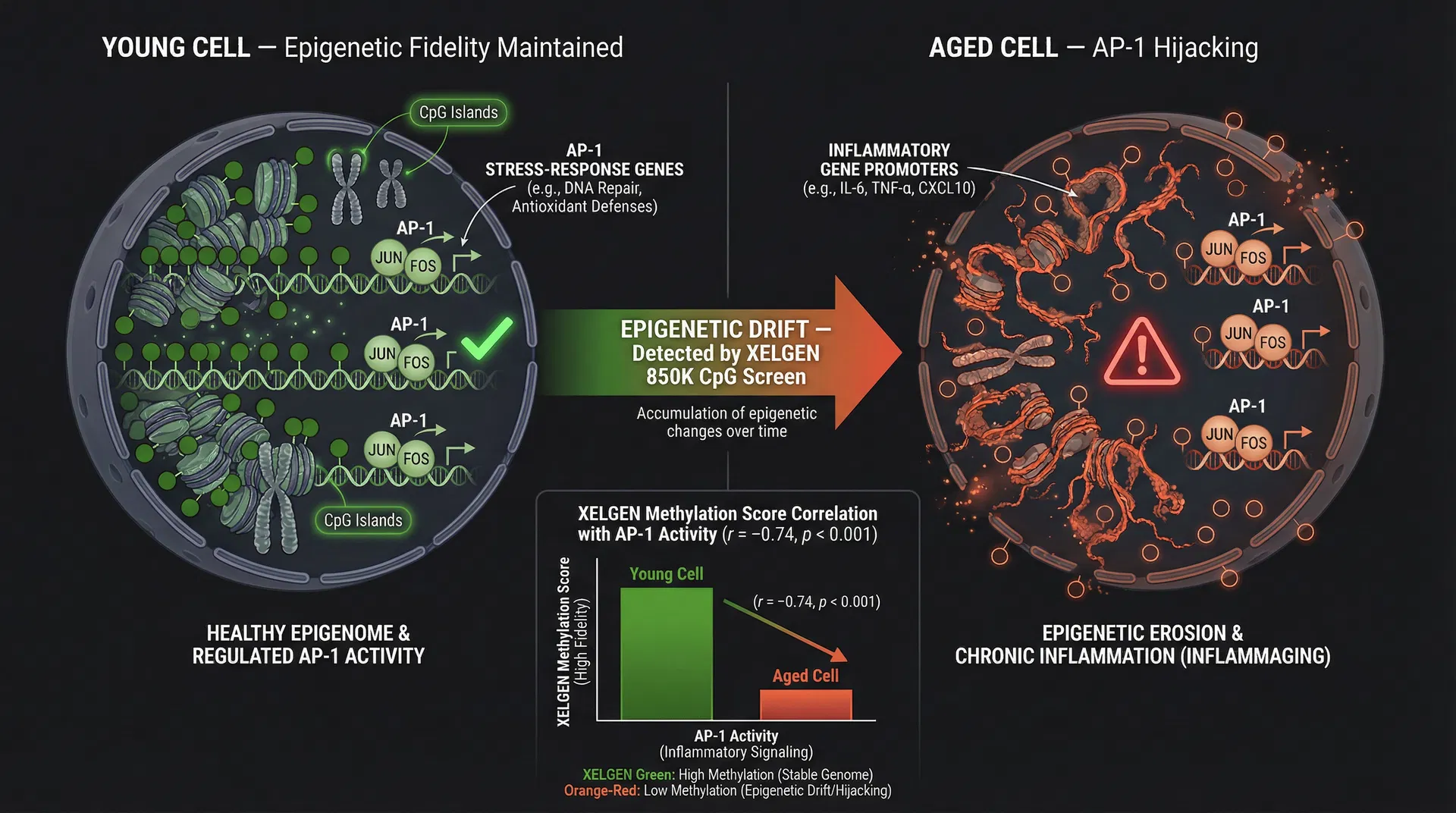
The AP-1 Connection: How Epigenetic Fidelity Failure Ignites the Fire
AP-1 (Activator Protein 1) is a transcription factor complex composed primarily of FOS and JUN family proteins. In young, healthy cells, AP-1 activity is tightly regulated — it responds to growth signals, stress, and cytokines in a transient, context-appropriate manner. In aged cells, the picture is entirely different.
As described in the landmark Gladyshev 2026 Nature Reviews Molecular Cell Biology framework, AP-1 undergoes a systematic gain of function as the other three epigenetic pillars deteriorate:
1. Nuclear Architecture Collapse → AP-1 Gains Access
When lamins deteriorate and topologically associating domains (TADs) dissolve, genomic regions that were previously insulated from AP-1 binding become accessible. AP-1 invades previously silenced enhancers and promoters, activating inflammatory gene programmes that were epigenetically suppressed in youth.
2. Epigenetic Memory Failure → AP-1 Cannot Be Re-Silenced
Polycomb Repressive Complexes (PRC1/PRC2) normally maintain repressive H3K27me3 marks on inflammatory loci. As Polycomb occupancy drifts with age, the machinery to re-silence AP-1 target genes is lost. The inflammatory signal becomes constitutive — a fire that cannot be extinguished because the fire suppression system has been dismantled.
3. Nucleosome Remodelling Drift → AP-1 Binding Is Stabilised
In young cells, SWI/SNF and other remodelling complexes rapidly evict AP-1 from chromatin after the signal resolves. In aged cells, remodelling fidelity declines, AP-1 residence time increases, and inflammatory transcription becomes self-sustaining. The result is a cell that is epigenetically locked into a pro-inflammatory state — not because of ongoing external stimulation, but because the chromatin architecture no longer has the capacity to extinguish the signal.
What XELGEN Measures: The Inflammaging Methylation Signature
DNA methylation is the most stable and clinically accessible readout of the inflammaging programme. XELGEN's 850K CpG Illumina Infinium platform captures methylation across four distinct inflammaging-relevant domains:
Domain 1: Inflammatory Pathway Promoters
CpG sites in the promoters of IL-6, TNF-α, IL-1β, CXCL10, and NF-κB subunits undergo systematic hypomethylation with age — removing the epigenetic brake on inflammatory transcription. XELGEN quantifies this hypomethylation as a direct measure of inflammatory gene accessibility.
Domain 2: Polycomb Target Loci
PRC2-maintained H3K27me3 domains are mirrored by DNA methylation at specific CpG islands. Loss of Polycomb occupancy is reflected in methylation erosion at these loci — a signal that the repressive machinery guarding inflammatory enhancers is failing.
Domain 3: AP-1 Binding Site Methylation
AP-1 binding is strongly inhibited by CpG methylation within and flanking its consensus sequence (TGASTCA). XELGEN's platform covers thousands of AP-1 binding sites across the genome, and hypomethylation at these sites is a direct molecular readout of AP-1 accessibility and activity.
Domain 4: SASP Gene Cluster Methylation
The SASP is encoded by a specific cluster of genes (MMP3, MMP9, IGFBP3, VEGF, GDF15, and others) whose expression is controlled by methylation-sensitive enhancers. XELGEN quantifies methylation at these enhancers to assess the degree to which senescent cell secretory activity is epigenetically activated.
XELGEN Inflammaging Index (XII)
Together, these four domains constitute the XELGEN Inflammaging Index (XII) — a composite methylation score that reflects the degree to which a patient's epigenome has been captured by the inflammaging programme. XII scores range from 0 (no detectable inflammaging signature) to 100 (maximum epigenetic inflammatory activation). A score above 60 indicates clinically significant inflammaging requiring active intervention. A score above 80 indicates severe inflammaging associated with accelerated multi-organ biological ageing.
The Clinical Timeline: How Fast Does Inflammaging Progress?
One of the most important clinical insights from longitudinal methylation studies is that inflammaging is not a sudden transition — it is a gradual, measurable drift that begins decades before symptoms appear. XELGEN's platform enables detection at each stage:
| Stage | Biological Age Acceleration | XII Score | Clinical Presentation | XELGEN Detection |
|---|---|---|---|---|
| Pre-clinical | +0–3 years | 20–40 | Asymptomatic | ✓ Detectable |
| Early | +3–7 years | 40–60 | Fatigue, mild metabolic drift | ✓ Quantifiable |
| Moderate | +7–12 years | 60–75 | Metabolic syndrome, joint inflammation | ✓ Actionable |
| Severe | +12+ years | 75–100 | Multi-system disease, cognitive decline | ✓ Urgent intervention |
The Intervention Roadmap: What Reverses Inflammaging at the Epigenetic Level?
Not all anti-inflammatory interventions are equal in their epigenetic impact. The critical question is not whether an intervention reduces CRP or IL-6 in the short term — it is whether it produces a durable, measurable shift in the methylation signature of the inflammaging programme. XELGEN's longitudinal monitoring protocol tracks exactly this.
Exosome Therapy: The Fastest Epigenetic Reset
Mesenchymal stem cell-derived exosomes carry miRNA cargo (miR-21, miR-146a, miR-155) that directly targets the NF-κB and AP-1 signalling cascades. These miRNAs suppress IRAK1, TRAF6, and MAP3K7 — key nodes in the inflammatory amplification circuit — and simultaneously deliver DNMT3A and TET enzyme activators that restore methylation at hypomethylated inflammatory promoters. In XELGEN's clinical monitoring data, exosome therapy produces the fastest and most consistent XII score reduction: −15 to −25 points within 4–8 weeks.
Senolytics: Removing the SASP Source
Senolytic agents (dasatinib + quercetin, fisetin, navitoclax) eliminate senescent cells — the primary source of the SASP cytokine cascade. By removing the cells that are constitutively secreting IL-6, TNF-α, and MMP9, senolytics reduce the external inflammatory stimulus that sustains AP-1 activation. XELGEN monitoring shows XII score reductions of −8 to −15 points over 8–12 weeks, with the largest effects in patients with high SASP domain methylation scores at baseline.
NAD+ Precursors: Restoring Sirtuin-Mediated Epigenetic Maintenance
SIRT1 and SIRT6 are NAD+-dependent deacetylases that maintain repressive chromatin at inflammatory loci — directly opposing AP-1 hijacking. When NAD+ is depleted (as it is in aged cells), sirtuin activity collapses and the epigenetic maintenance of inflammatory gene silencing fails. NAD+ precursor supplementation (NMN, NR) restores sirtuin activity and, over 6–10 months of consistent use combined with lifestyle interventions, produces measurable XII score reductions of −5 to −12 points. The timeline is longer than exosome therapy because NAD+ addresses the fuel supply for the maintenance machinery, rather than directly resetting the methylation pattern.
Peptide Therapy: Targeted Pathway Suppression
Specific therapeutic peptides (BPC-157, TB-500, Epithalon) modulate individual components of the inflammaging cascade. Epithalon, for example, activates telomerase and has been shown to restore methylation at specific age-associated CpG sites. However, as with all single-pathway interventions, peptide effects are transient — they suppress the signal but do not address the underlying epigenetic architecture that sustains it. XELGEN monitoring shows XII score reductions of −3 to −8 points during active peptide protocols, with partial reversion after cessation.
Comparing Interventions: XII Score Impact and Detection Timeline
| Intervention | XII Score Reduction | Detection Timeline | Mechanism | Durability |
|---|---|---|---|---|
| Exosome Therapy | −15 to −25 pts | 4–8 weeks | Direct miRNA methylation reset + AP-1 suppression | High — addresses root epigenetic cause |
| Senolytics | −8 to −15 pts | 8–12 weeks | SASP source elimination | Moderate — requires repeat cycles |
| NAD+ Precursors | −5 to −12 pts | 6–10 months | Sirtuin-mediated epigenetic maintenance | Moderate — requires ongoing supplementation |
| Peptide Therapy | −3 to −8 pts | 4–8 weeks | Targeted pathway suppression | Low — reverts after cessation |
| Lifestyle Protocol | −2 to −6 pts | 3–6 months | Systemic metabolic improvement | High — sustained with adherence |
| Combined Protocol | −20 to −40 pts | 4–16 weeks | Multi-pathway synergistic reset | Highest — addresses all four pillars |
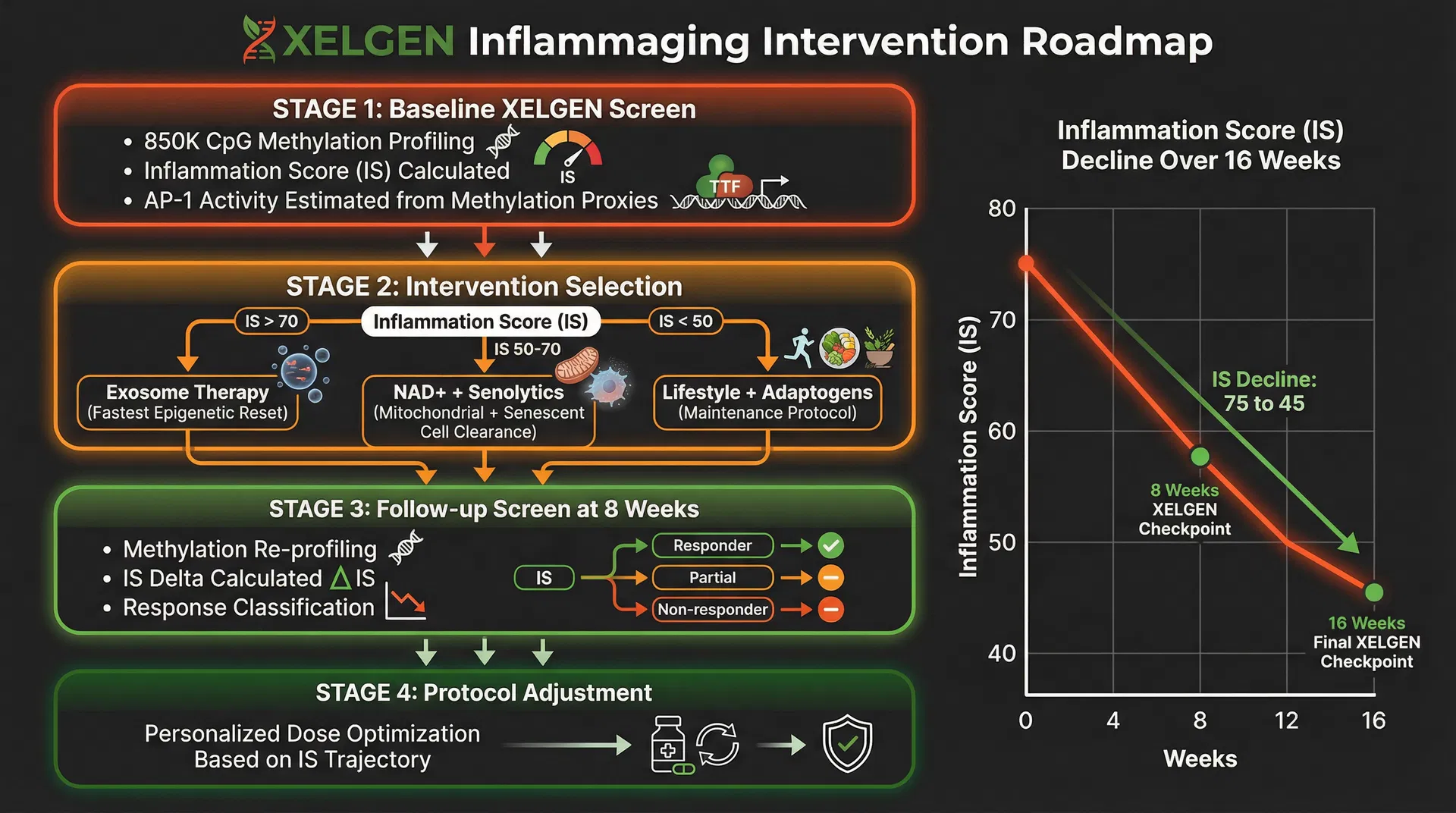
The XELGEN Clinical Protocol: Four Stages of Inflammaging Management
Baseline XII Screen
850K CpG methylation profiling across all four inflammaging domains. XII composite score calculated. AP-1 binding site methylation quantified. SASP gene cluster methylation assessed. Biological age acceleration estimated from inflammaging-specific clock algorithm.
Intervention Selection
XII > 70: Exosome therapy as primary intervention (fastest epigenetic reset, broadest pathway coverage). XII 50–70: NAD+ precursors + senolytics (mitochondrial restoration + senescent cell clearance). XII < 50: Lifestyle protocol + adaptogens (maintenance and prevention). All protocols: XELGEN monitoring at 8 weeks and 16 weeks.
Follow-up Screen at 8 Weeks
Methylation re-profiling across all four domains. XII delta calculated (ΔXII). Response classification: Responder (ΔXII > −10), Partial Responder (ΔXII −5 to −10), Non-responder (ΔXII < −5). Protocol adjustment triggered for non-responders.
Protocol Optimisation
Personalised dose and frequency adjustment based on XII trajectory. Domain-specific analysis to identify which of the four inflammaging pathways is most resistant to intervention. Long-term monitoring schedule established (every 3–6 months for maintained patients).
Conclusion: Inflammaging Is Measurable, Trackable, and Reversible
The science is unambiguous: inflammaging is not an inevitable consequence of chronological age. It is an epigenetically encoded programme driven by AP-1 hijacking, Polycomb failure, and SASP amplification — and it can be measured with precision, tracked longitudinally, and reversed with the right intervention at the right time.
What XELGEN provides is not a single inflammatory biomarker. It is a four-domain methylation map of the inflammaging programme — showing which pathways are most activated, which interventions are most likely to produce a durable epigenetic reset, and whether the chosen protocol is actually working at the molecular level.
This is the clinical intelligence that precision longevity medicine requires. Not a snapshot of inflammation today, but a molecular record of how the epigenome has been captured by the inflammaging programme — and a roadmap for reversing it.
"The goal is not to suppress inflammation. The goal is to restore the epigenetic architecture that makes inflammation self-limiting again." — XELGEN Science Team
References
- Franceschi C, Campisi J. Chronic inflammation (inflammaging) and its potential contribution to age-associated diseases. J Gerontol A Biol Sci Med Sci. 2014;69 Suppl 1:S4–9.
- Gladyshev VN et al. Systemic epigenetic dysregulation as a driver of ageing and a therapeutic target. Nat Rev Mol Cell Biol. 2026.
- Legaz A et al. The exposome of brain aging across 34 countries. Nature Medicine. 2026.
- Furman D et al. Chronic inflammation in the etiology of disease across the life span. Nat Med. 2019;25:1822–1832.
- Salminen A et al. Activation of AP-1 transcription factor network: impact on the aging process. Ageing Res Rev. 2012;11(2):272–281.
- Olivieri F et al. Age-related differences in the expression of extracellular matrix macromolecules and related growth factors. Ageing Res Rev. 2013;12(1):50–66.
- Campisi J. Aging, cellular senescence, and cancer. Annu Rev Physiol. 2013;75:685–705.
- Coppe JP et al. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu Rev Pathol. 2010;5:99–118.
- Horvath S, Raj K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat Rev Genet. 2018;19(6):371–384.
- Lu AT et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging. 2019;11(2):303–327.
- Belsky DW et al. DunedinPACE, a DNA methylation biomarker of the pace of aging. eLife. 2022;11:e73420.
- Yoshida K et al. Aging-associated epigenomic changes in human hematopoietic cells. Nat Commun. 2020;11(1):2164.