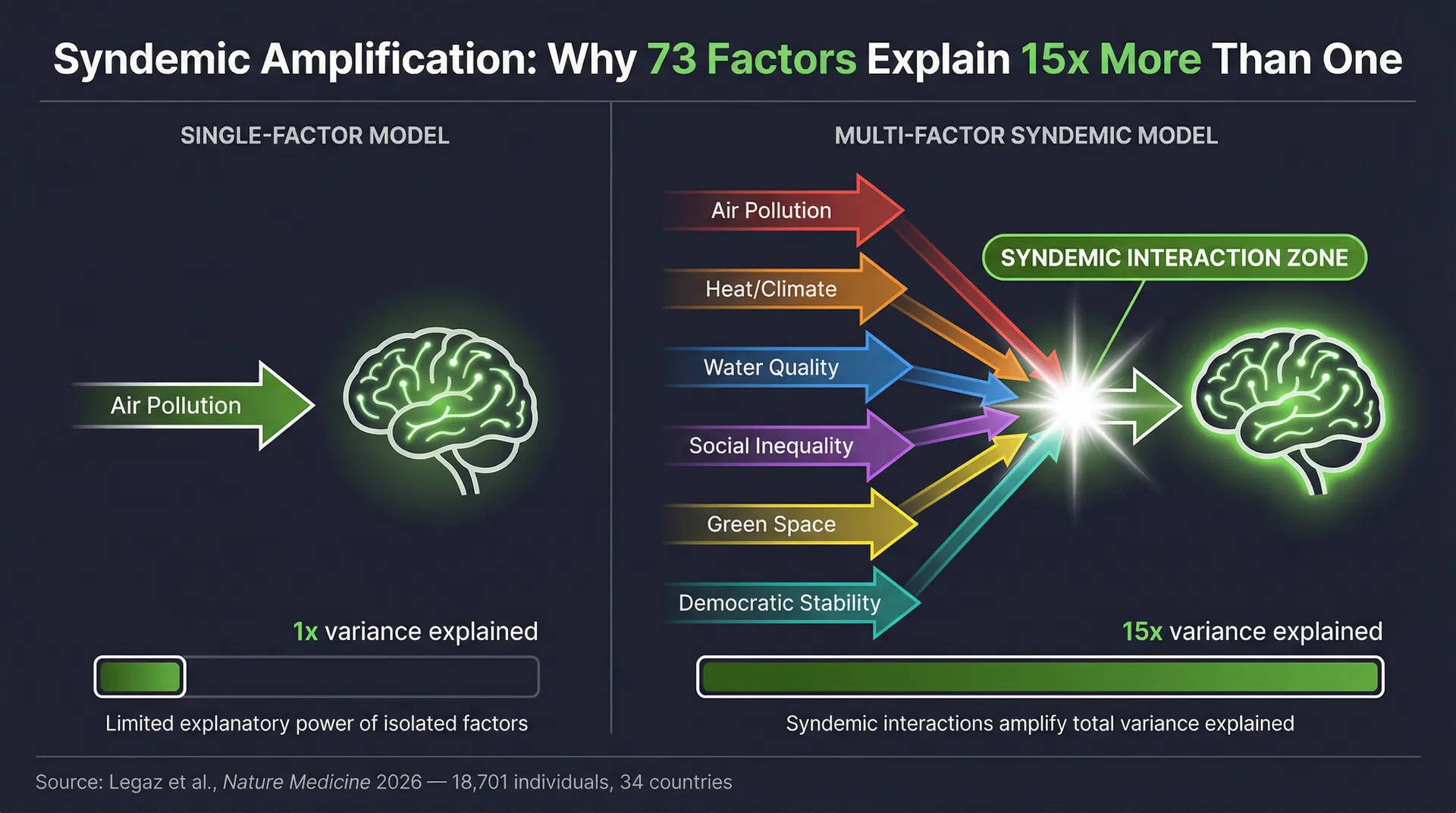
A landmark study published in Nature Medicine on 3 April 2026 — 18,701 individuals across 34 countries, 73 environmental factors — has delivered a quantitative verdict on one of the most important questions in biological aging science: when you model multiple exposures together, they explain up to 15 times more variance in brain aging than any single exposure alone. The effects are syndemic and nonlinear. Interactions across domains amplify biological impact in ways that no single-factor model can detect, predict, or address. This is not a finding about the brain alone. It is a finding about the fundamental nature of biological aging — and it has direct implications for every clinical platform that claims to measure or reverse it.
This article examines what the Nature Medicine exposome study found, why its central conclusion — that multi-factor interaction is the dominant driver of biological age acceleration — validates the architectural logic of XELGEN's 850K CpG methylation platform, and what it means for physicians building precision longevity protocols for their patients.
1. The Single-Factor Trap in Aging Medicine
For decades, the dominant clinical approach to biological aging has been reductionist. Measure one biomarker. Target one pathway. Prescribe one intervention. Telomere length. CRP. Fasting glucose. NAD+ levels. Each of these has genuine biological relevance. But each tells only a partial story — and none, in isolation, can capture the true complexity of how the human body ages.
The problem is not that these biomarkers are wrong. It is that they are structurally incomplete. Biological aging is not a single-pathway phenomenon. It is the cumulative, interactive result of decades of environmental exposures — physical, social, metabolic, psychological — each leaving molecular traces in the epigenome, each modifying the cellular response to every other exposure. A patient with high CRP and high cortisol does not simply have two independent problems. The inflammatory and stress-axis pathways interact, amplify each other, and together produce a biological age acceleration that is greater than the sum of either alone.
This is precisely what the Nature Medicine study has now demonstrated at global scale.
2. What the Nature Medicine Exposome Study Found
The study, led by Agustín Ibáñez and colleagues at the Global Brain Health Institute, Trinity College Dublin, analysed 18,701 individuals across 34 countries spanning six continents. Using neuroimaging-derived brain age gap (BAG) as the primary outcome — the difference between estimated brain biological age and chronological age — the researchers modelled 73 environmental indicators across two broad domains.
| Exposure Domain | Key Indicators | Primary Brain Structures Affected | Biological Mechanism |
|---|---|---|---|
| Physical Exposome | Air pollution, extreme heat, green space access, water quality | Hippocampus, amygdala, brainstem autonomic centres | Neuroinflammation, oxidative stress, vascular dysfunction |
| Social Exposome | Socioeconomic inequality, democratic stability, social support | Prefrontal cortex, anterior cingulate, limbic system | Chronic HPA axis activation, allostatic load, neuroplasticity impairment |
The critical finding was not that each domain independently accelerated brain aging — that was already known. The critical finding was the syndemic interaction effect: when both physical and social exposome burdens co-occur — as they almost always do in real patients — the combined biological impact is not additive. It is multiplicative. Chronic stress amplifies the neuroinflammatory response to air pollution. Poverty reduces access to green space and healthcare, compounding the vascular effects of climate extremes. Social isolation amplifies the cortisol-mediated effects of socioeconomic precarity. The exposures interact, and their interactions accelerate biological aging far beyond what any single-factor model can predict.
"We aimed to test whether the combined syndemic effects of environmental exposures could better explain variation in brain aging across populations than individual exposures or single clinical diagnoses."
— Agustín Ibáñez, Principal Investigator, Global Brain Health Institute, Trinity College Dublin
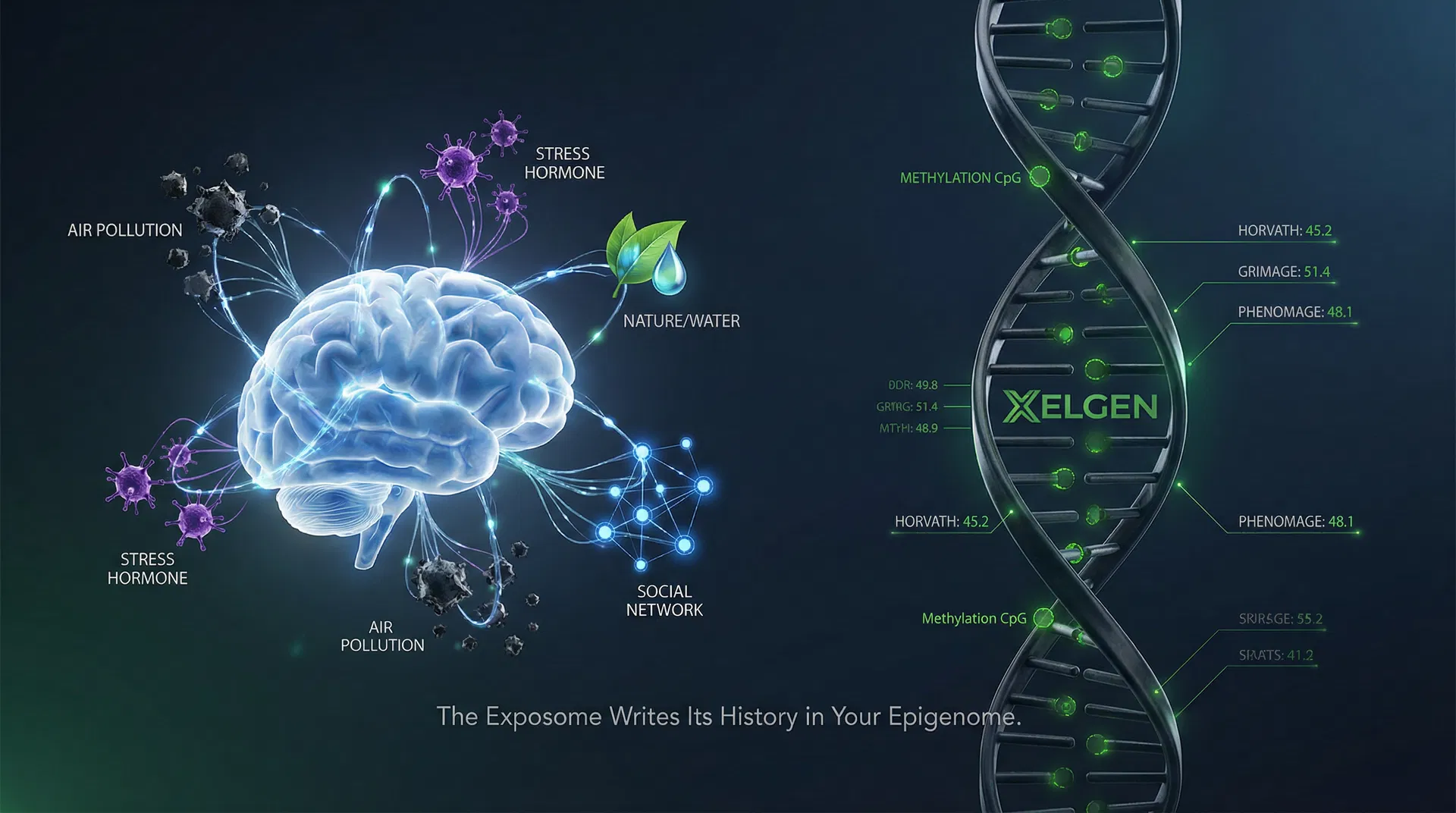
3. Where the Exposome Writes Its History: The Methylome
The exposome is the cumulative collection of all environmental, social, and physiological exposures an individual experiences from conception to the present. It encompasses everything the genome does not encode: the air breathed, the food consumed, the stress experienced, the social connections maintained or lost, the pollutants absorbed, the sleep quality maintained or degraded over decades.
The critical question for clinical medicine is: where in the body does the exposome leave its trace? The answer, established by two decades of epigenetics research, is the DNA methylome. Every significant environmental exposure — chronic stress, air pollution, dietary quality, physical activity, toxic chemical exposure, social isolation — modifies the methylation state of cytosine residues at CpG dinucleotides across the genome. These modifications accumulate, compound, and interact. And they are measurable.
What the Nature Medicine study adds is the quantitative proof that this cumulative multi-factor signal is not merely additive — it is multiplicatively amplified by interactions between exposure domains. The epigenome does not record each exposure separately. It records the integrated, interactive result of all exposures simultaneously. This is precisely why genome-wide methylation analysis — not targeted single-gene panels, not single-pathway biomarkers — is the appropriate clinical tool for biological age assessment.
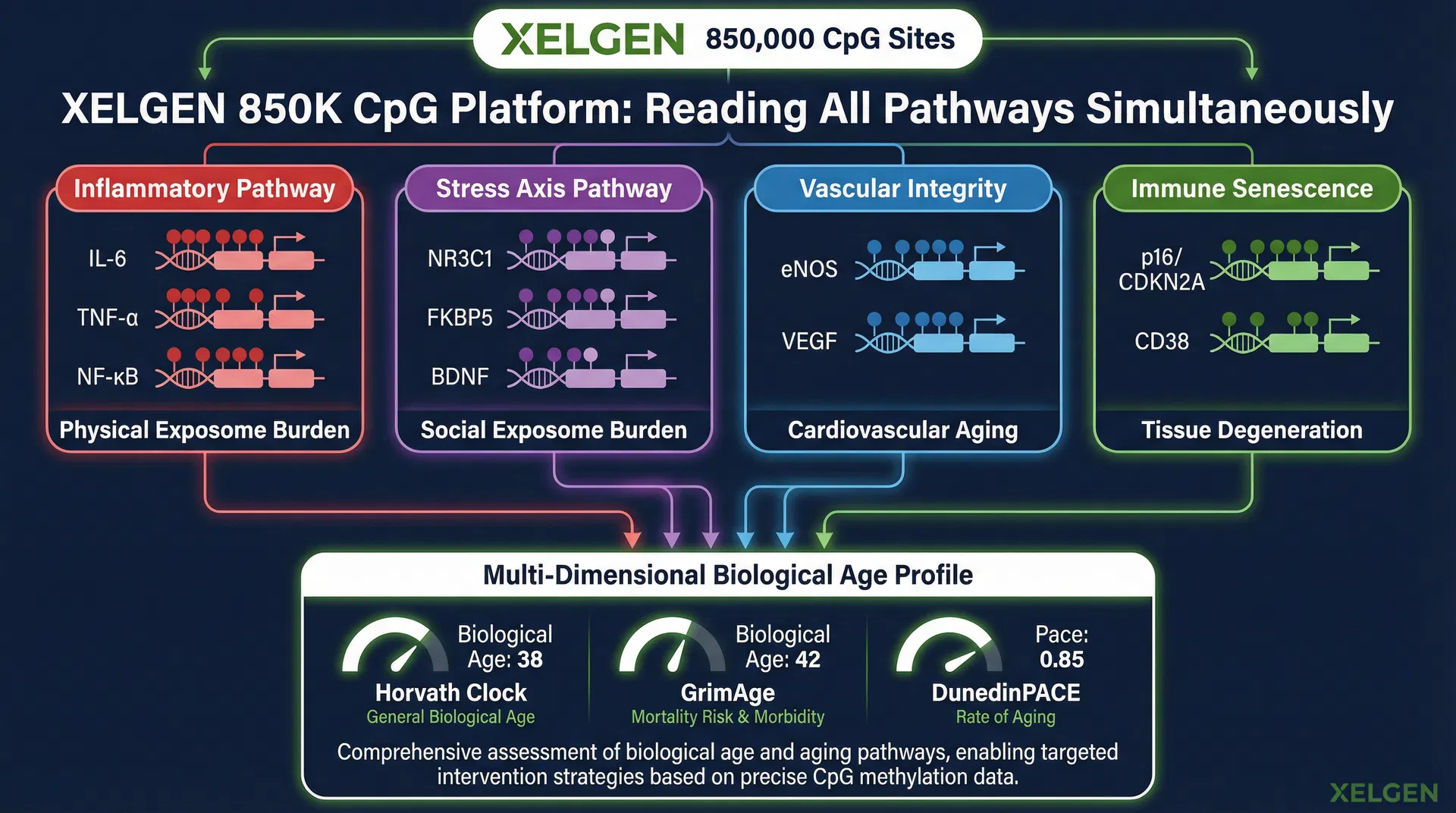
4. How XELGEN's 850K Platform Reads All Pathways Simultaneously
XELGEN's Illumina Infinium EPIC array analyses 850,000 CpG sites across the human genome — the most comprehensive commercial methylation platform available. This breadth is not incidental. It is the architectural requirement for capturing the multi-factor syndemic signal that the Nature Medicine study has now quantified at global scale.
The four pathway domains that XELGEN's platform reads simultaneously map directly onto the two exposome domains identified in the Nature Medicine study:
| XELGEN Pathway | Key Gene Loci | Exposome Domain Captured | Clinical Significance |
|---|---|---|---|
| Inflammatory Pathway | IL-6, TNF-α, NF-κB, CRP | Physical exposome burden | Cumulative neuroinflammatory and systemic inflammatory load |
| Stress Axis Pathway | NR3C1, FKBP5, BDNF, SLC6A4 | Social exposome burden | Cumulative HPA axis activation, allostatic load, neuroplasticity impairment |
| Vascular Integrity | eNOS, VEGF, endothelial loci | Physical + metabolic exposome | Cardiovascular aging, cerebrovascular health, tissue perfusion |
| Immune Senescence | p16/CDKN2A, CD38, SASP genes | Combined exposome burden | Pace of immunological aging, SASP-driven tissue degeneration |
Critically, XELGEN does not report a single biological age number. It reports a multi-dimensional biological age profile using three validated clock algorithms simultaneously — Horvath (general biological age), GrimAge (mortality risk and morbidity), and DunedinPACE (current rate of aging). Each clock is sensitive to different pathway combinations. Together, they provide a clinical picture that no single algorithm can produce alone — mirroring the multi-factor modelling approach that the Nature Medicine study has now validated at population scale.
5. From Population Science to Individual Patient Intelligence
The Nature Medicine study operates at the country level. It tells us that populations exposed to higher combined physical and social exposome burdens age faster biologically. But it cannot tell a clinician what is happening in the specific patient sitting in the consultation room.
This is where XELGEN's individual-level methylation screen becomes the clinical translation of population science. A patient's 850K CpG profile encodes their personal exposome history — not a country's average air quality, but the specific inflammatory, stress-axis, and vascular burden that this patient's cells have accumulated over a lifetime. XELGEN does not infer biological age from population averages. It reads the molecular record that the patient's own cells have written.
The XELGEN Multi-Factor Clinical Protocol
Multi-Factor Baseline Assessment
The XELGEN screen establishes the patient's current biological age across multiple clock algorithms, identifies which exposure pathways are most activated, and quantifies the gap between biological and chronological age. Unlike single-biomarker assessments, this baseline captures the syndemic interaction pattern — which pathways are co-activated and amplifying each other.
Precision Intervention Targeting
Because XELGEN identifies which methylation pathways are most dysregulated, physicians can direct interventions precisely: exosome therapy for patients showing accelerated immune senescence and vascular aging; stress-axis interventions for patients with NR3C1 and FKBP5 hypermethylation; anti-inflammatory protocols for patients with IL-6 and TNF-α pathway activation.
Objective Response Monitoring
Repeat XELGEN screening at 3–6 month intervals provides the only objective indicator of whether an intervention has produced epigenetic change — not just symptomatic improvement, but measurable reversal of biological age acceleration at the molecular level.
6. The Multi-Factor Imperative
The Nature Medicine exposome study is a turning point in biological aging science. Its central finding — that multi-factor interaction explains up to 15× more variance in brain aging than single exposures — is not a finding about the brain alone. It is a finding about the fundamental nature of biological aging: it is inherently multi-factorial, nonlinear, and syndemic.
Single biomarkers, single clocks, and single-pathway interventions will always be limited by this reality. The clinical tool that matches the complexity of the problem is a genome-wide, multi-pathway, multi-clock epigenomic platform — one that reads all accumulated exposures simultaneously, identifies the interaction patterns driving biological age acceleration, and gives physicians the intelligence to intervene precisely and monitor objectively.
This is what XELGEN Cell Intelligence was built to do. The science has now confirmed, at global scale, why it matters.
Key Takeaways for Clinicians
| Finding | Clinical Implication |
|---|---|
| 73 factors jointly explain 15× more variance than one | Single-biomarker aging assessment is structurally insufficient |
| Physical and social exposomes affect different brain regions via different mechanisms | Multi-pathway assessment is required for intervention targeting |
| Syndemic effects are nonlinear and interaction-dependent | No single clock or biomarker can capture the full aging signal |
| XELGEN 850K CpG screen reads all pathways simultaneously | Delivers the multi-factor intelligence the science demands |
| Epigenetic response to intervention detectable at 4–6 weeks (exosomes) | XELGEN monitoring enables objective, timely assessment of treatment efficacy |
References
- Legaz A, Moguilner S, Hernández H, et al. The exposome of brain aging across 34 countries. Nature Medicine. 2026. DOI: 10.1038/s41591-026-04302-z
- Horvath S. DNA methylation age of human tissues and cell types. Genome Biology. 2013;14(10):R115.
- Lu AT, et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging. 2019;11(2):303–327.
- Belsky DW, et al. DunedinPACE, a DNA methylation biomarker of the pace of aging. eLife. 2022;11:e73420.
- Baccarelli AA, Woychik RJ. Epigenetics and the exposome. NIEHS Environmental Factor. February 2025.
- Gladyshev VN. Systemic epigenetic dysregulation as a driver of ageing. Nature Reviews Molecular Cell Biology. 2026.