Table of Contents
- 1Biological Aging as a Systems Process
- 2Chronological Age vs Biological Age
- 3DNA Methylation as a Biomarker of Aging
- 4Genome-Scale DNA Methylation Profiling
- 5Technology Platform: Illumina Infinium Arrays
- 6Comparison of DNA Methylation Technologies
- 7Quality Control and Analytical Rigor
- 8XELGEN Cell Intelligence Systems Biology Framework
- 9Longitudinal Biological Aging Trajectories
- 10Illumina Probe Architecture
- 11CpG Genomic Coverage
- 12Epigenetic Clock Training
- 13XELGEN System Interaction Heatmap
- 14Conclusion
- 15References
Aging is the dominant risk factor for most chronic diseases, including cardiovascular disease, neurodegeneration, metabolic disorders, and cancer. However, chronological age alone does not accurately reflect physiological aging or health status.
The concept of biological age has emerged as a powerful framework for understanding individual aging trajectories. Among biological aging biomarkers, DNA methylation has emerged as one of the most robust and widely validated indicators of biological age. Epigenetic clocks derived from methylation patterns have demonstrated strong correlations with chronological age, disease risk, and mortality outcomes.
The XELGEN platform integrates genome-scale DNA methylation profiling with advanced bioinformatics to evaluate epigenetic biomarkers and systems-level biological aging processes. This white paper describes the biological foundations, technology platform, analytical rigor, and clinical applications of the XELGEN Cell Intelligence Platform.
Biological Aging as a Systems Process
Aging is a multifactorial biological process characterized by progressive functional decline across multiple physiological systems. Research has identified interconnected mechanisms driving aging, including genomic instability, epigenetic alterations, mitochondrial dysfunction, cellular senescence, stem cell exhaustion, and chronic inflammation.
Key Hallmarks of Aging
These processes influence tissue regeneration, immune response, metabolic regulation, and cognitive function. Importantly, individuals age at different rates. Environmental exposures, lifestyle behaviors, and genetic predispositions influence biological aging trajectories. Understanding these trajectories requires molecular biomarkers capable of capturing aging processes directly.
Chronological Age vs Biological Age
Chronological age measures the passage of time. Biological age reflects physiological aging and health status. Individuals with accelerated biological aging often exhibit increased disease risk and reduced physiological resilience.
The distinction between chronological and biological age is clinically significant. Two patients of identical chronological age may have biological ages differing by a decade or more, reflecting fundamentally different disease risk profiles, treatment responses, and longevity trajectories. The XELGEN platform quantifies this difference with genome-scale precision.
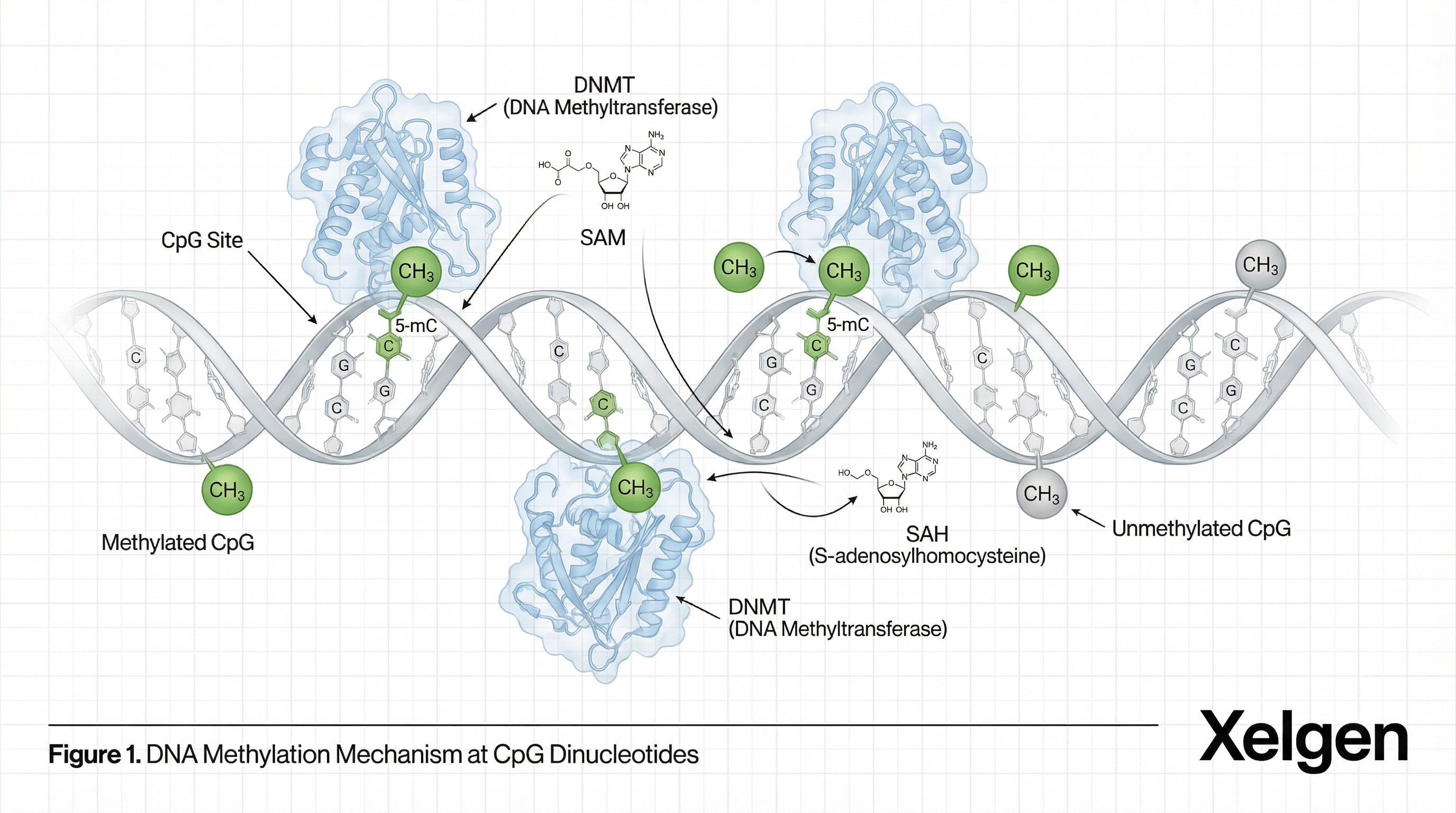
DNA Methylation as a Biomarker of Aging
DNA methylation is an epigenetic modification involving the addition of methyl groups to cytosine bases within CpG dinucleotides. These modifications regulate gene expression without altering DNA sequence and play important roles in gene regulation, cellular differentiation, development, and aging processes.
Systematic changes in CpG methylation occur during aging, enabling the development of epigenetic clocks. These clocks estimate biological age based on methylation patterns across selected CpG loci. The Illumina Infinium EPIC BeadChip interrogates more than 850,000 CpG sites, providing the genome-scale resolution necessary for robust biological age estimation.
Validated Epigenetic Clocks Used by XELGEN
Multi-tissue epigenetic clock trained on 353 CpG sites across 51 tissue types. Correlation r=0.96 with chronological age.
Blood-based epigenetic clock using 71 CpG markers. Strong predictor of all-cause mortality.
Phenotypic age clock integrating clinical biomarkers. Predicts morbidity, mortality, and physiological resilience.
Composite mortality predictor integrating DNAm-based surrogates of plasma proteins. Strongest predictor of lifespan.
Genome-Scale DNA Methylation Profiling
Genome-wide methylation profiling enables measurement of methylation patterns across hundreds of thousands of CpG sites simultaneously. This approach supports epigenome-wide association studies (EWAS), aging biomarker discovery, and disease risk analysis at a resolution impossible with targeted or reduced-representation methods.
The XELGEN platform leverages genome-scale profiling to capture the full complexity of the epigenetic aging landscape. Rather than relying on a limited panel of CpG sites, XELGEN's analysis encompasses the regulatory regions, enhancers, promoters, and gene bodies that collectively define biological age with clinical-grade precision.
Technology Platform: Illumina Infinium DNA Methylation Arrays
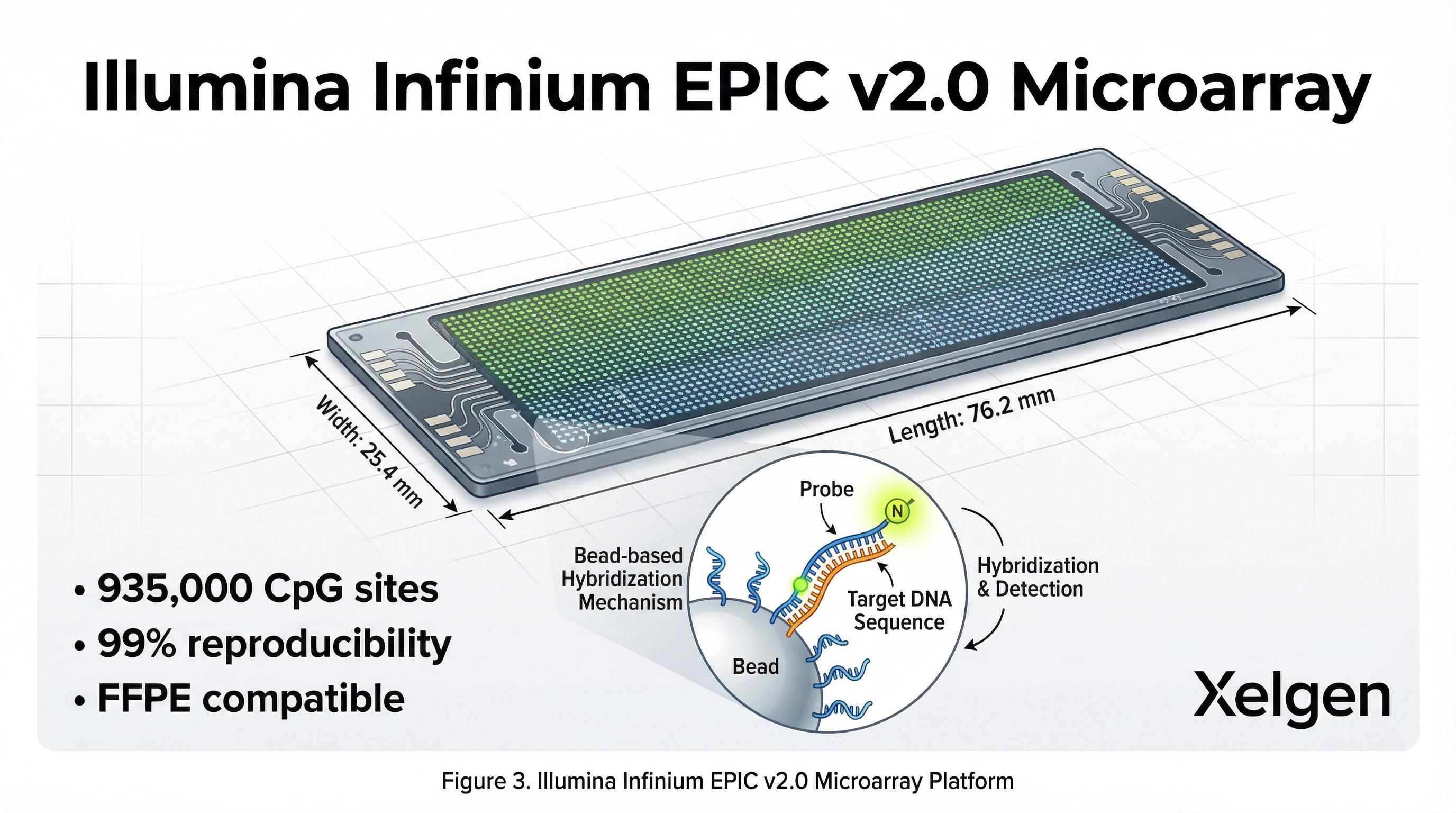
The XELGEN platform utilizes the Illumina Infinium methylation assay, one of the most widely used and validated technologies for genome-wide methylation profiling. The Infinium HumanMethylationEPIC BeadChip interrogates more than 850,000 CpG sites across regulatory regions of the genome.
Key technical specifications of the Illumina EPIC array as deployed in the XELGEN platform:
| Specification | Value |
|---|---|
| CpG Sites Interrogated | >850,000 |
| Genomic Coverage | Promoters, enhancers, CpG islands, gene bodies, open sea regions |
| Technology | Infinium I & II bead-based assay |
| Sample Input | 250–500 ng genomic DNA |
| Sample Type | Whole blood, PBMC, saliva, FFPE tissue |
| Workflow Duration | ~2 weeks (sample to report) |
| Reproducibility | >99% concordance between technical replicates |
| Bisulfite Conversion Efficiency | >95% required for QC pass |
| Clinical Validation | CLIA-certified laboratory processing |
Comparison of DNA Methylation Technologies
Multiple technologies exist for methylation profiling. The choice of platform significantly impacts CpG coverage, reproducibility, cost, and clinical scalability. XELGEN's selection of the Infinium EPIC array reflects its optimal balance of genome-wide coverage, validated performance, and clinical throughput.
| Parameter | WGBS | RRBS | Targeted Seq. | Infinium EPIC |
|---|---|---|---|---|
| CpG Coverage | ~28M | ~3M | Custom | >850K |
| Genome Coverage | Complete | CpG-rich regions | Targeted loci | Regulatory regions |
| Cost per Sample | $$$$ | $$$ | $$ | $$ |
| Throughput | Low | Medium | Medium | High |
| Reproducibility | High | Medium | High | Very High |
| Clinical Scalability | Limited | Moderate | Moderate | Excellent |
| Validation for Aging Clocks | Limited | Limited | Limited | Extensive |
Quality Control and Analytical Rigor
Reliable biomarker analysis requires robust quality control procedures at every stage of the analytical pipeline. The XELGEN platform implements a three-tier QC framework:
Sample QC
- DNA integrity assessment (DIN ≥7)
- Bisulfite conversion efficiency (>95%)
- DNA concentration verification
- Sample identity confirmation
Array QC
- Probe detection p-values (<0.01)
- Signal intensity metrics
- Background fluorescence correction
- Staining and hybridization controls
Bioinformatic QC
- BMIQ normalization algorithm
- ComBat batch effect correction
- Outlier sample detection
- Cross-array concordance validation
XELGEN Cell Intelligence Systems Biology Framework
Most biomarker platforms analyze isolated measurements or simple timepoint comparisons. The XELGEN Cell Intelligence framework interprets genome-scale methylation data through a systems biology perspective, evaluating interactions across biological systems including immune regulation, metabolic pathways, mitochondrial function, and cellular stress responses.
This systems-level interpretation enables XELGEN to deliver insights that extend beyond a single biological age number. The platform identifies which biological systems are aging at accelerated rates, which are well-preserved, and how these systems interact — providing clinicians with actionable intelligence for precision longevity medicine protocols.
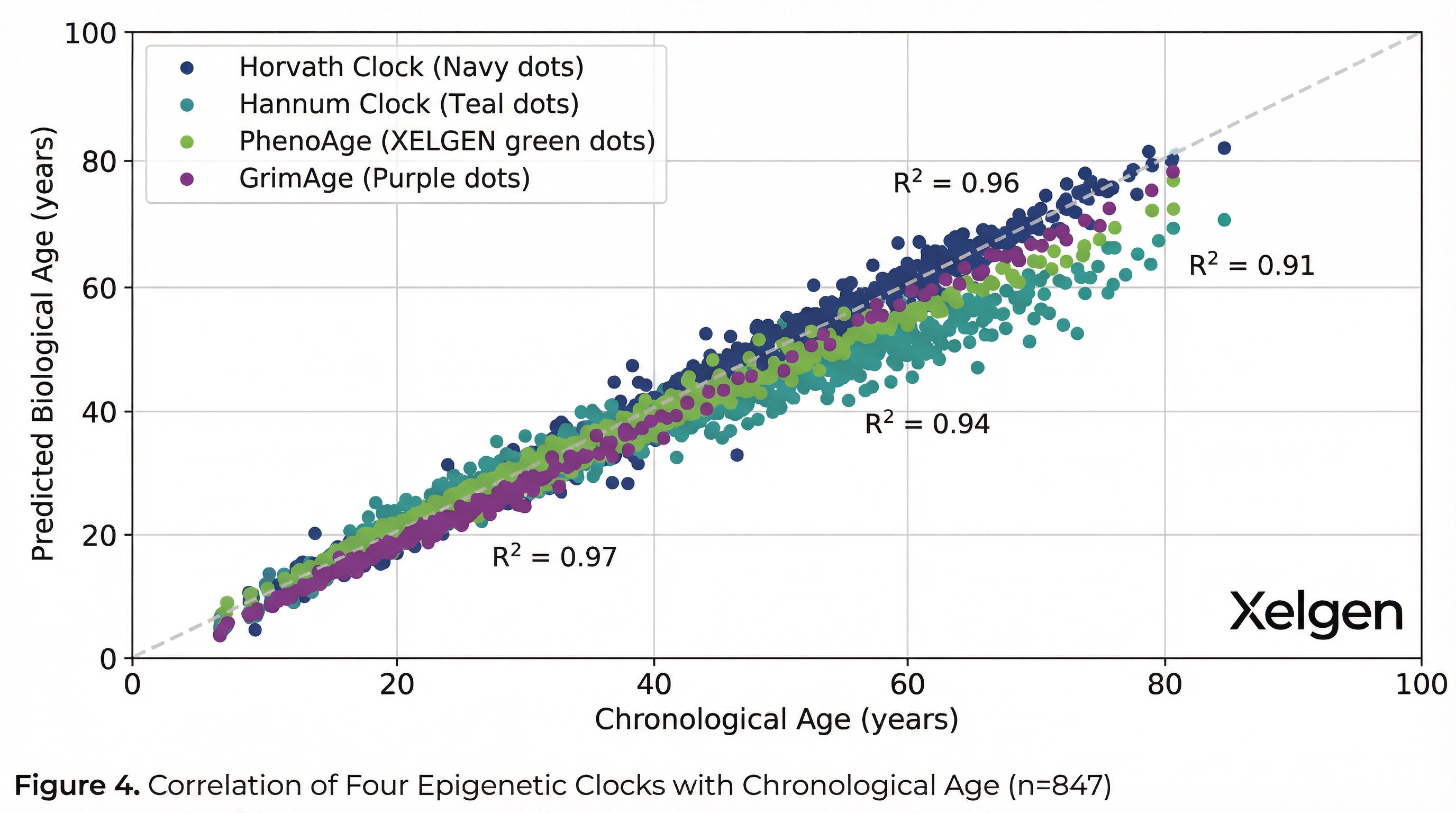
Longitudinal Biological Aging Trajectories
Biological aging is dynamic and influenced by lifestyle, disease, and environmental exposures. Longitudinal monitoring enables evaluation of lifestyle interventions, metabolic therapies, and regenerative medicine treatments. The XELGEN platform supports serial testing to track biological age trajectories over time.
Clinical evidence from longitudinal methylation studies demonstrates that biological age can be meaningfully modified. Interventions including caloric restriction, exercise protocols, and regenerative therapies have demonstrated measurable reductions in epigenetic age acceleration. XELGEN's longitudinal tracking capability provides the quantitative framework for evaluating these interventions in clinical practice.
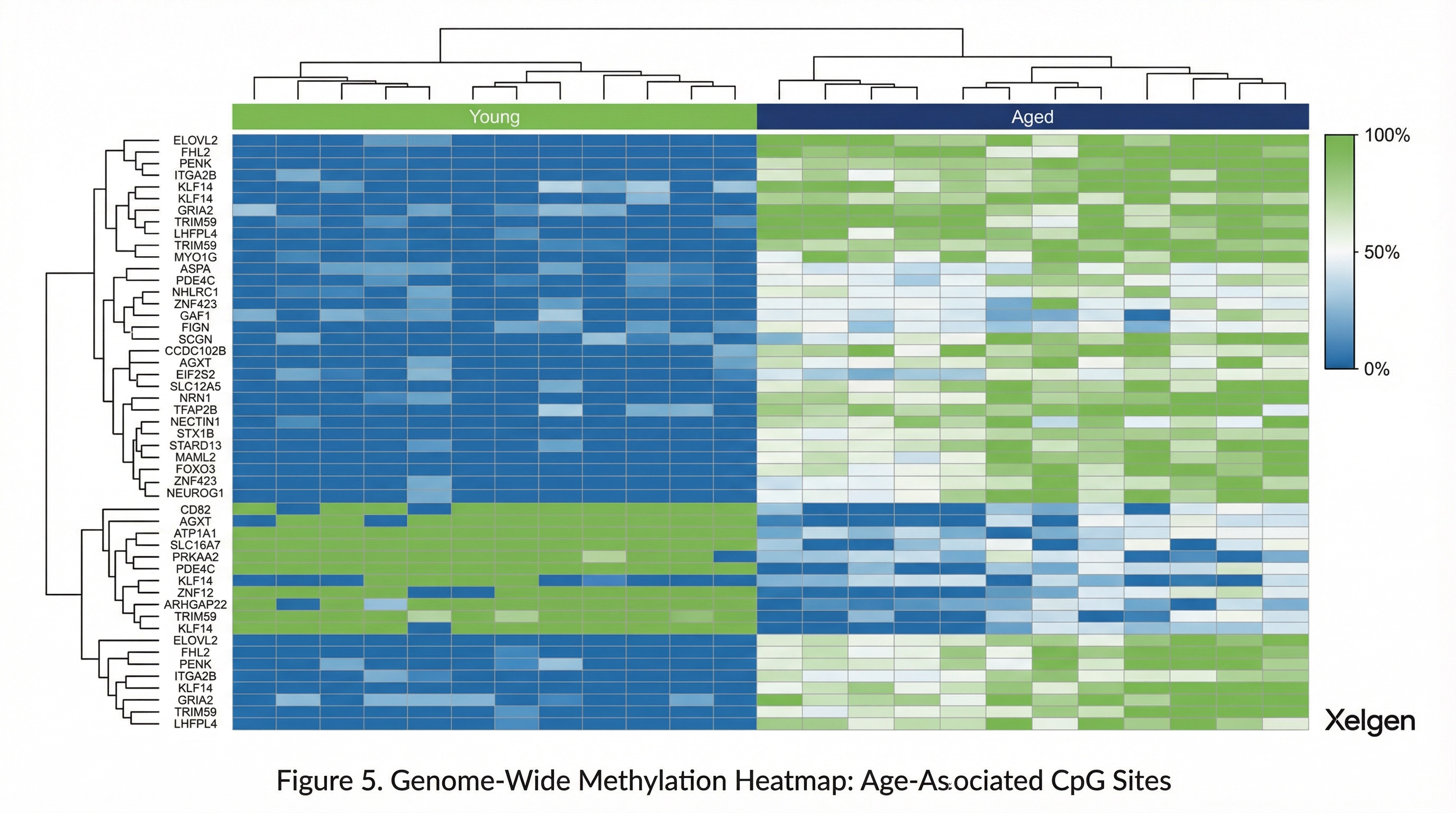
Illumina Probe Architecture
Infinium arrays measure methylation using bead-bound probes targeting CpG loci. Two probe chemistries are employed: Infinium Type I probes (two bead types per CpG, higher sensitivity) and Infinium Type II probes (single bead type per CpG, higher throughput). The EPIC array uses both chemistries optimized for their respective genomic contexts.
Each probe undergoes single-base extension following hybridization to bisulfite-converted DNA. Fluorescent signal intensities are captured and converted to beta values (0 = unmethylated, 1 = fully methylated) for each CpG site, providing a continuous quantitative measure of methylation level.
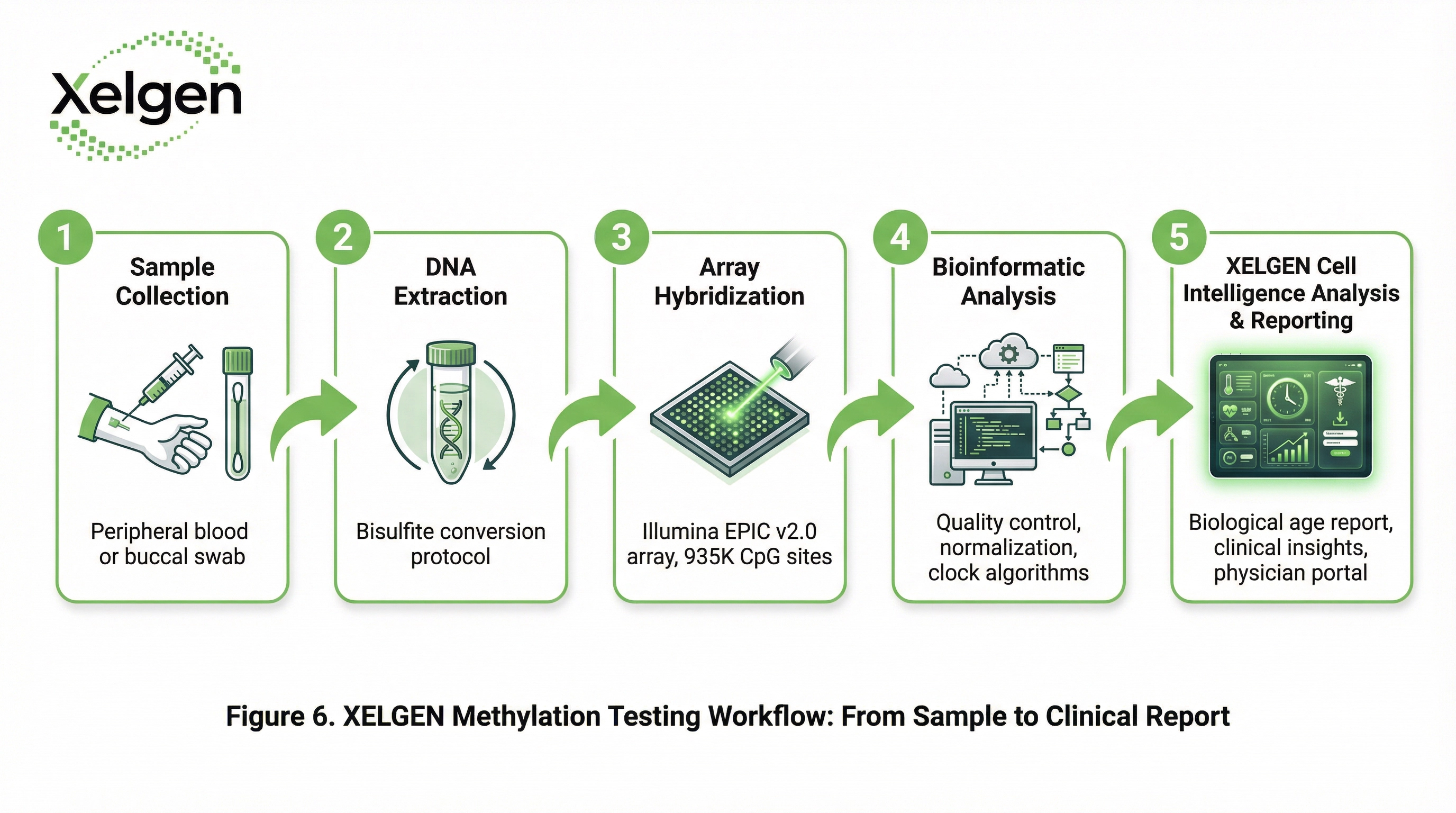
CpG Genomic Coverage
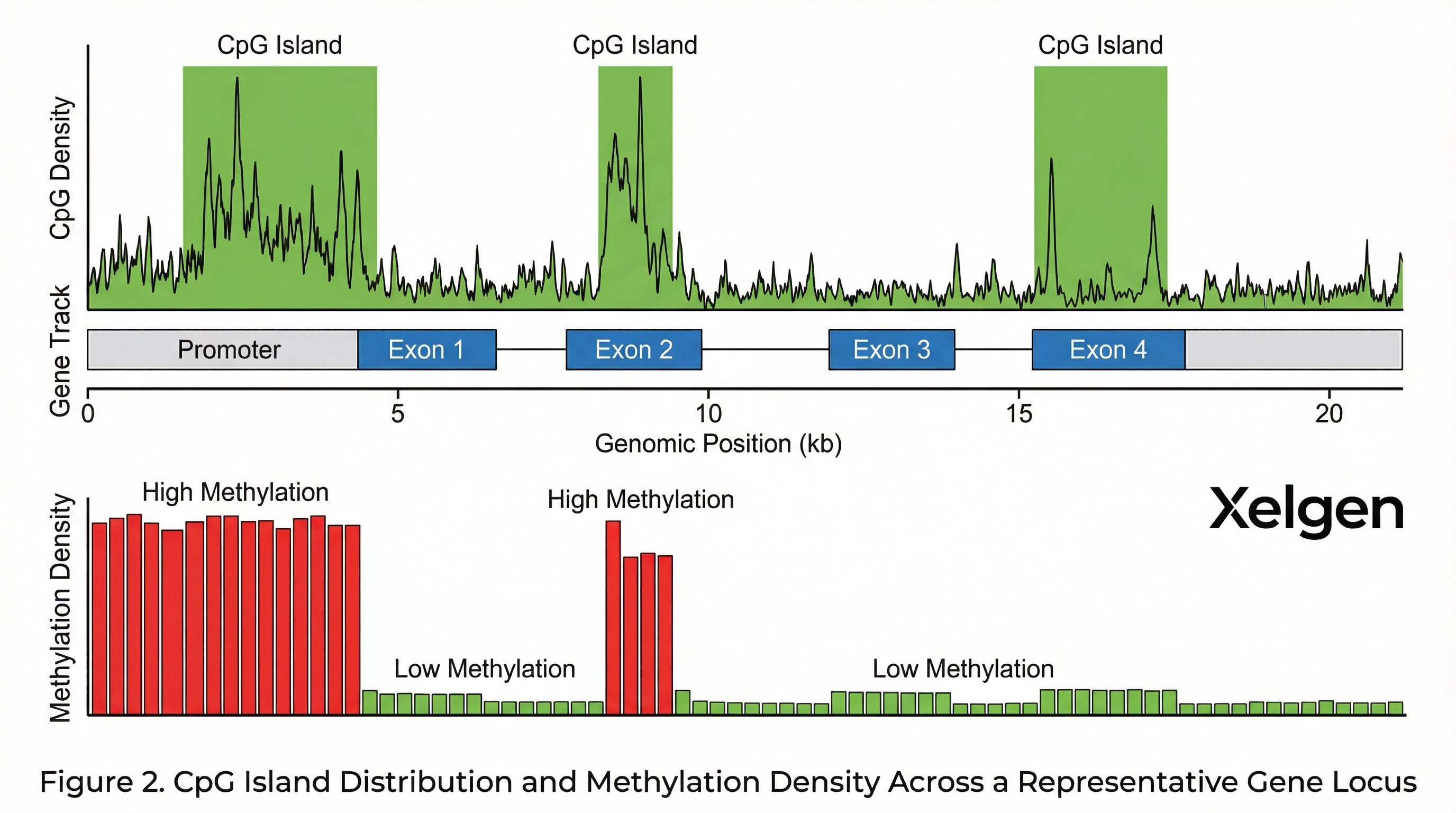
Genome-wide arrays target CpG sites across regulatory regions. The Infinium EPIC array covers CpG sites in promoters, enhancers, CpG islands, gene bodies, and open sea regions — providing comprehensive coverage of the regulatory epigenome. This breadth of coverage is essential for capturing the full spectrum of aging-associated methylation changes.
Approximately 3% of the human genome consists of CpG dinucleotides, but these are not uniformly distributed. CpG islands — regions of high CpG density typically associated with gene promoters — are particularly important for aging clock analysis. The EPIC array provides dense coverage of these functionally critical regions.
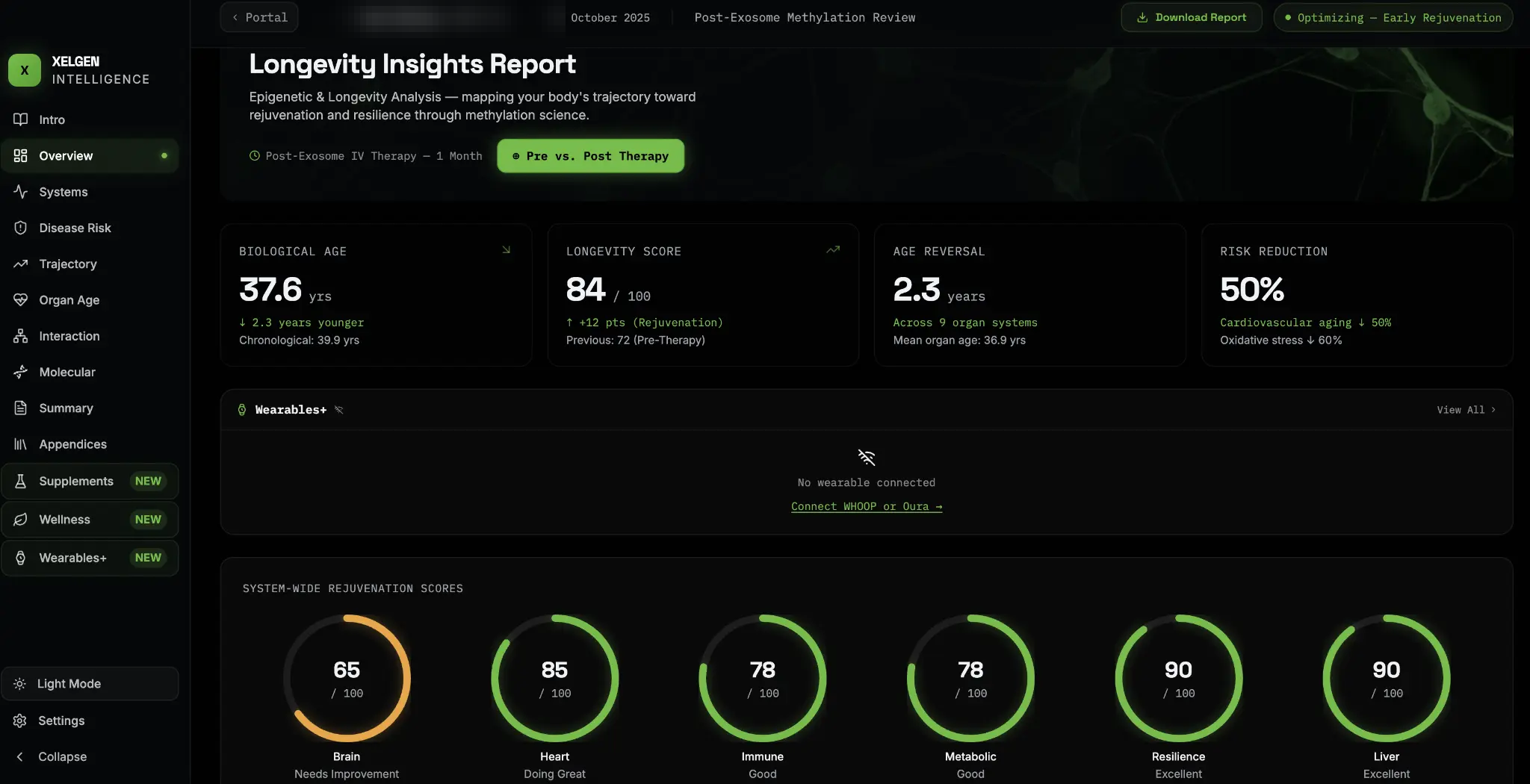
Epigenetic Clock Training
Epigenetic clocks are developed using machine learning models trained on large methylation datasets. The training process involves feature selection from hundreds of thousands of CpG sites, elastic net regression to identify the most informative loci, and validation across independent cohorts spanning multiple tissue types and disease states.
The clocks deployed in the XELGEN platform have been validated in studies encompassing tens of thousands of individuals, with cross-cohort reproducibility exceeding r=0.95 for chronological age prediction. GrimAge, the most prognostically powerful clock, was trained on plasma protein surrogates and demonstrates significant associations with time-to-death independent of chronological age.
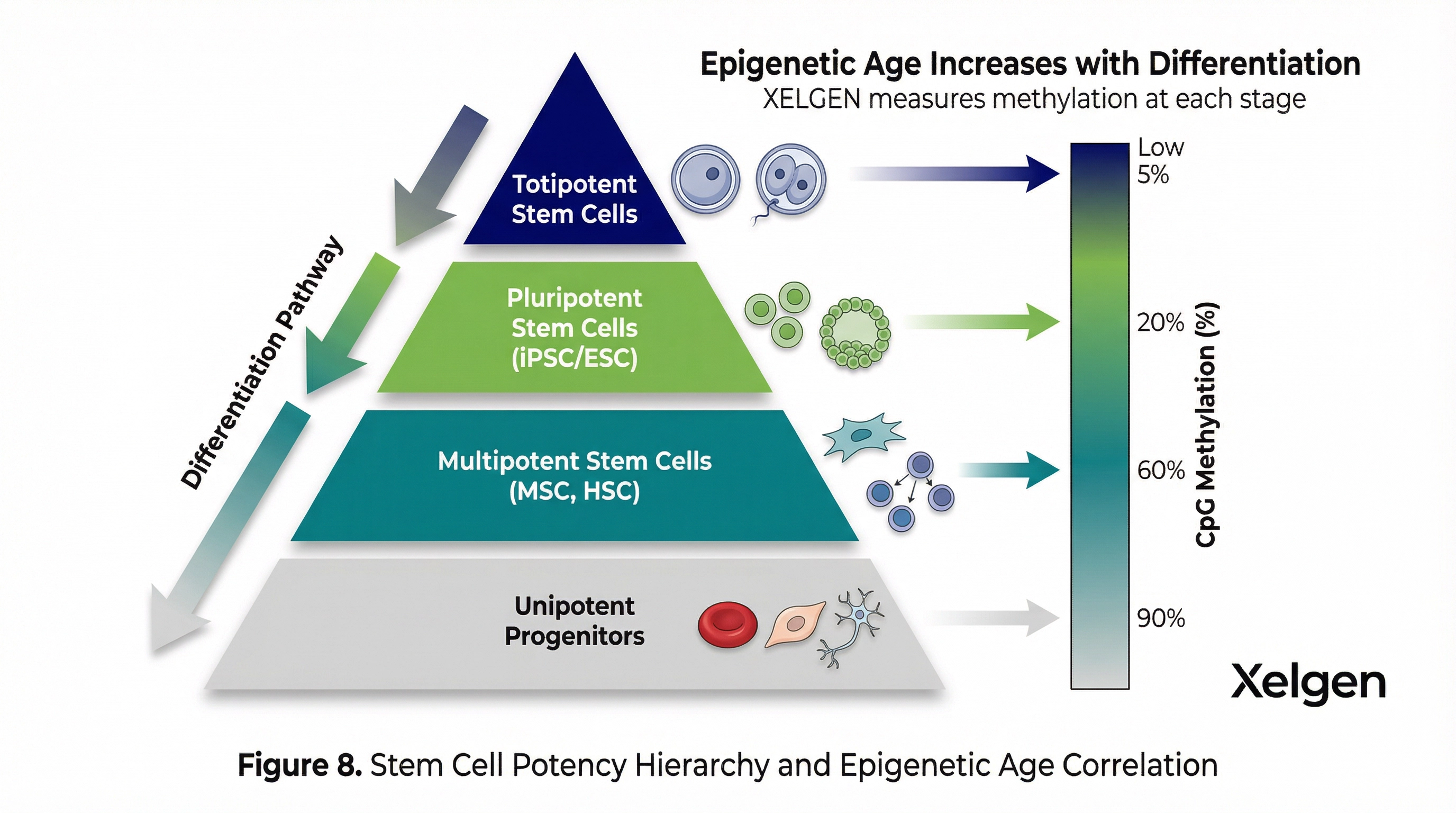
XELGEN System Interaction Heatmap
Genome-scale methylation data enables analysis of interactions between biological systems. The XELGEN Cell Intelligence platform generates a systems interaction map that quantifies the degree of epigenetic dysregulation across and between biological systems — immune function, metabolism, mitochondrial integrity, cellular stress, senescence burden, and DNA repair capacity.
This interaction heatmap provides clinicians with a visual representation of where biological aging is most advanced and which systems are most interconnected in a given patient's aging profile. This information directly informs the selection and sequencing of longevity medicine interventions.
Conclusion
Biological age biomarkers provide a powerful framework for understanding physiological aging processes and predicting health outcomes. The XELGEN platform integrates genome-scale DNA methylation profiling, rigorous analytical pipelines, and systems-level biological interpretation through the XELGEN Cell Intelligence framework.
This integrated approach enables advanced biomarker-driven insights into human aging and supports the emerging field of precision longevity medicine. By measuring biological age with genome-scale precision, XELGEN provides clinicians with the quantitative foundation necessary to evaluate, monitor, and optimize regenerative medicine interventions.
The XELGEN Cell Intelligence Platform represents a significant advancement in the clinical application of epigenetic biomarkers — moving biological age from a research metric to a clinically actionable tool for regenerative medicine and longevity medicine practice.
References
Seminal multi-tissue epigenetic clock using 353 CpG sites.
Development and validation of the PhenoAge clock.
GrimAge: strongest epigenetic predictor of mortality.
Technical validation of the EPIC BeadChip platform.
Comprehensive review of epigenetic clock methodology.
AI integration with epigenetic biomarker platforms.
Download the Full White Paper
Access the complete scientific overview with all 9 figures, methodology details, and clinical integration guidance.
